Publications
Peer Reviewed (most recent ones)
most updated list on google scholar
5) J.C. Geue, P. Liu, S. Keobouasone, P. Wilson, M. Manseau (2025).MhGeneS: An analytical pipeline to allow for robust microhaplotype genotyping. Molecular Ecology Resources; DOI:10.1111/1755-0998.14027
4) S.Hoban, I. Paz-Vinas, R.E. Shaw, L. Castillo-Reina, J.M. da Silva, J. A. DeWoody, R. Ekblom, A. Fedorca, B.R. Forester, W. C. Funk, J.C. Geue, M. Heuertz, P.M. Hollingsworth, A.C. Hughes, M.E. Hunter, C. Hvilsom, F. Ishihama, R. Jordan, B. Kalamujic Stroil, F. Kershaw, C.K. Khoury, V. Köppä, L. Laikre, A..J. MacDonald, A. Mastretta-Yanes, M.H. Meek, J. Mergeay, K.L. Milette, D. O'Brien, V.J. Rincon-Parra, M. A. Rodriguez-Morales, M.C. Schuman, G. Segelbacher, P. Sunnucks, R. S. Taylor, H. Thurfjell, C. Vernesi, C.E. Grueber (2024) DNA-based studies and genetic indicator assessments are complementary approaches to conserving evolutionary potential. Conservation Genetics; DOI: 10.1007/s10592-024-01632-8.
3) H.Hinneberg, T. Bamann, J.C. Geue, K. Foerster, A. Kupfer, H. A. Thomassen (2022). Truly invasive or simply non-native? Insights from an artificial crested newt hybrid zone. Conservation Science and Practice; DOI:10.1111/csp2.12752
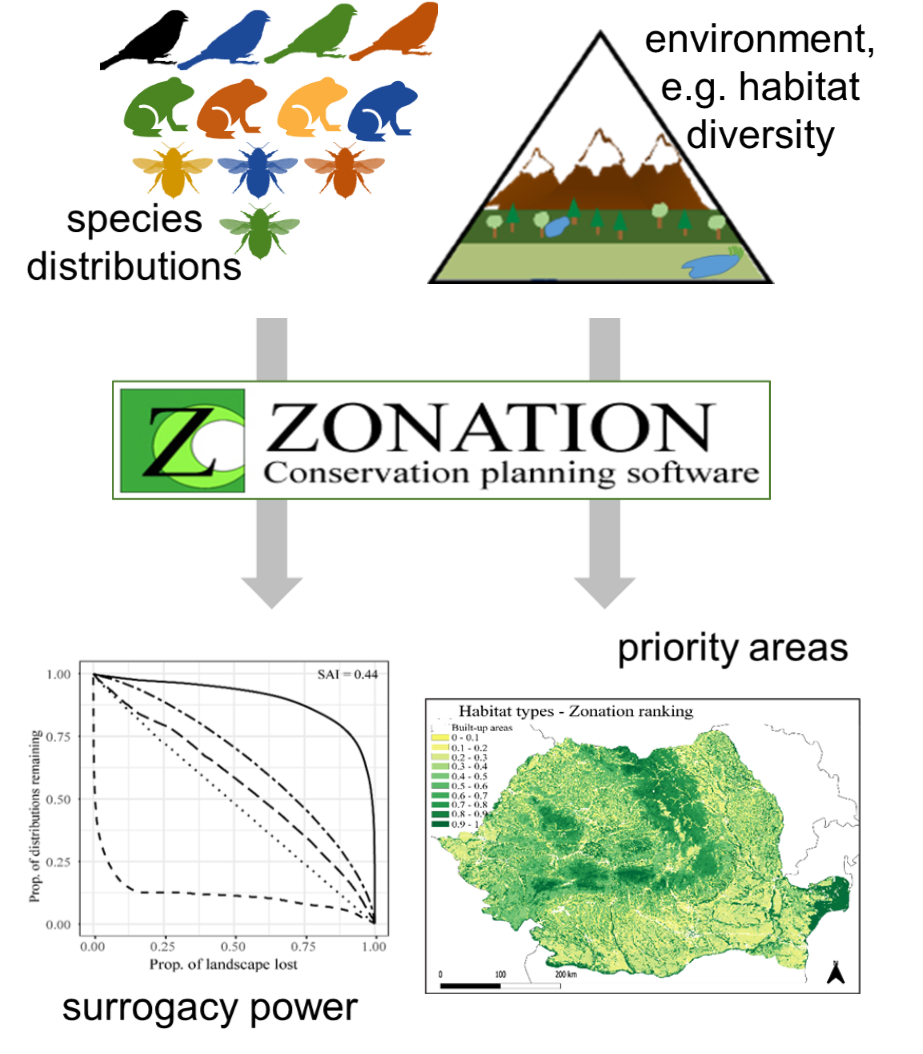
2) J.C. Geue, P.J. Rotter, C. Gross, Z. Benkő, I. Kovács, C. Fântână, J. Veres-Szászka,C. Domșa, E. Baltag, S.J. Daróczi, G.M. Bóné, V.D. Popescu, H.A. Thomassen (2022). Limited reciprocal surrogacy of bird and habitat diversity and inconsistencies in their representation in Romanian protected areas. PLOS ONE; DOI:10.1371/journal.pone.0251950
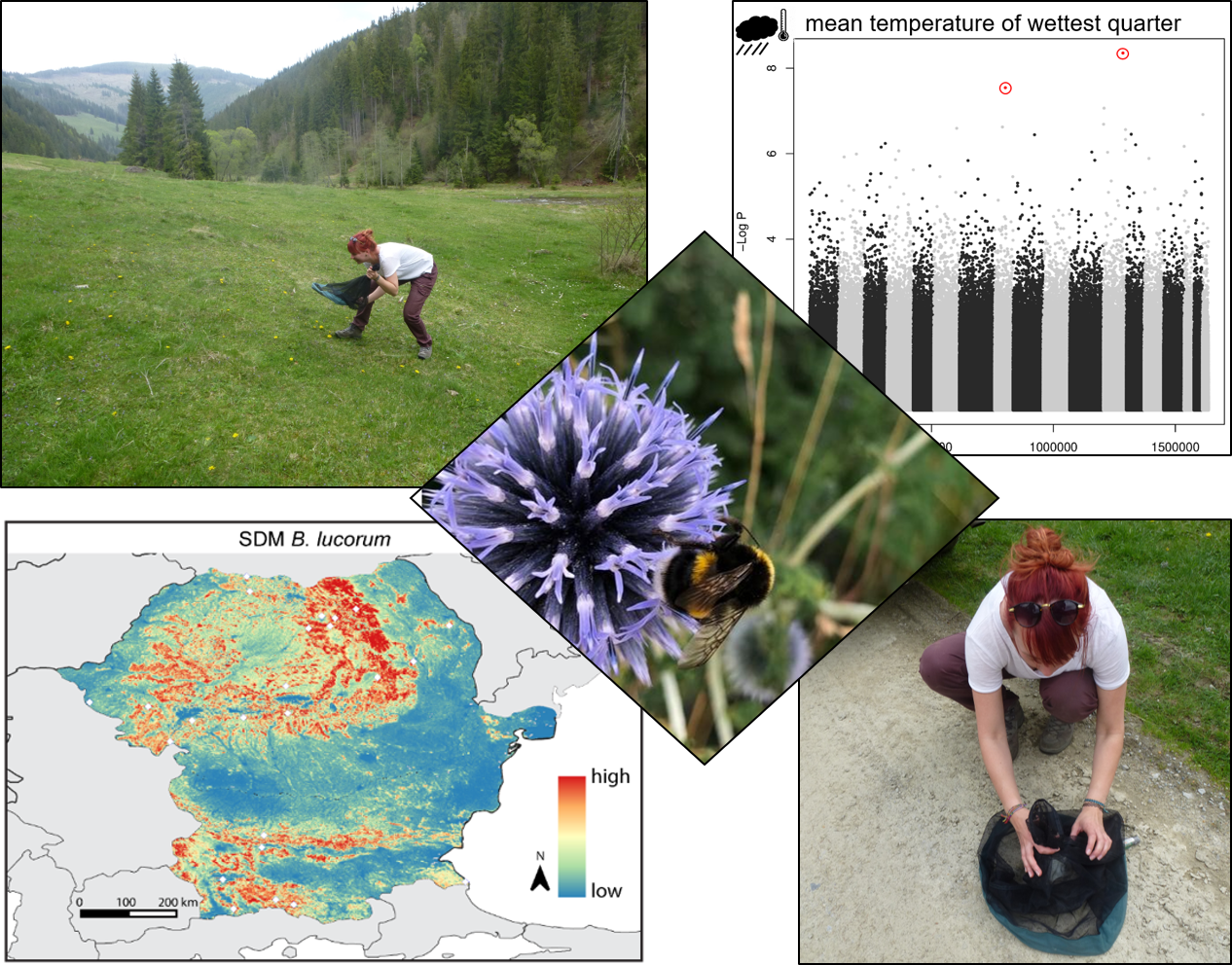
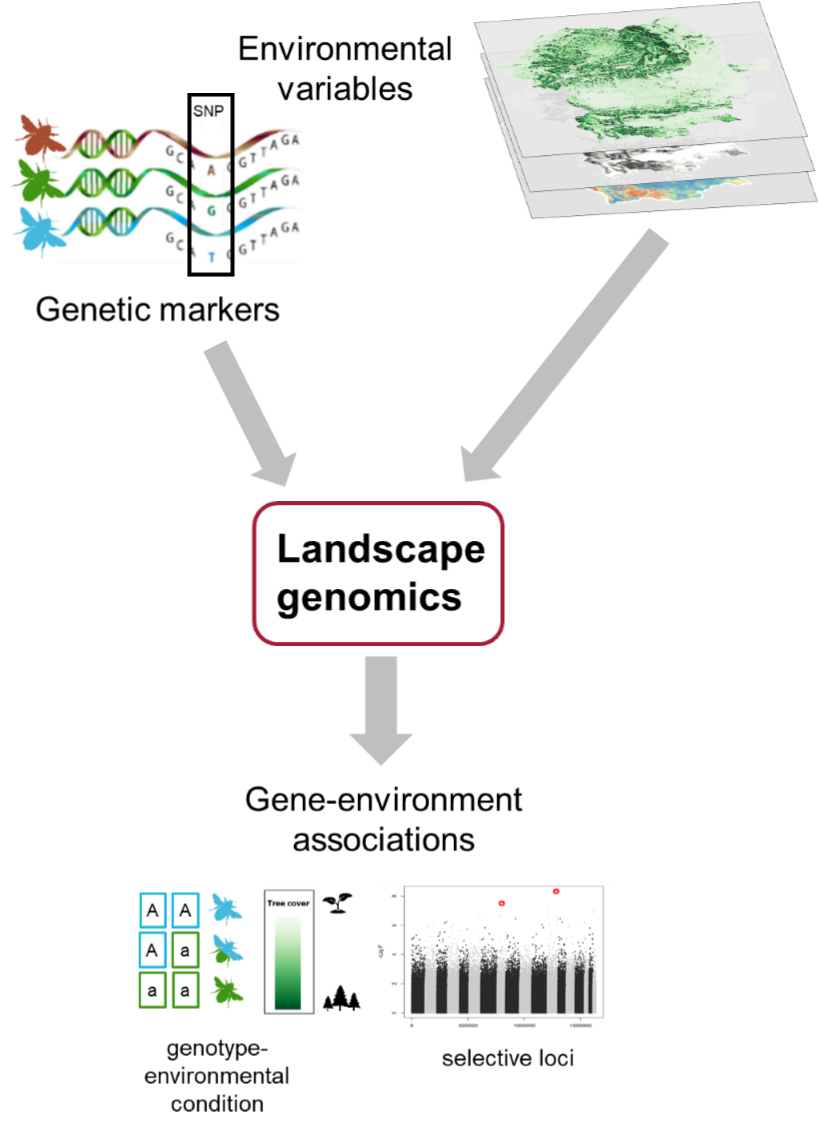
1) M.Glück, J.C. Geue, H.A. Thomassen (2022). Environmental differences explain subtle yet detectable genetic structure in a widespread pollinator. BMC Ecology; DOI: 10.1186/s12862-022-01963-5
Outreach
1) J.C. Geue and H.A. Thomassen (2021). Biodiversität: Mehr als nur
Artenvielfalt. IDM-Themenhefts Info Europa; PDF(in german)